Use user defined pseudotime¶
CAPITAL can now also use pseudotime that are calculated in other methods.
[1]:
import capital as cp
import scanpy as sc
import numpy as np
In this tutorial, we will use the same datasets as previous tutorial.
[2]:
adata1 = cp.dataset.setty19()
The dataset already exist in ../data/capital_dataset/setty19_capital.h5ad.
Reading datasets from ../data/capital_dataset/setty19_capital.h5ad.
[3]:
adata1
[3]:
AnnData object with n_obs × n_vars = 5780 × 1999
obs: 'n_genes', 'leiden'
var: 'n_cells', 'highly_variable', 'means', 'dispersions', 'dispersions_norm'
uns: 'diffmap_evals', 'hvg', 'leiden', 'leiden_colors', 'leiden_sizes', 'neighbors', 'paga', 'pca', 'umap'
obsm: 'X_diffmap', 'X_pca', 'X_umap'
varm: 'PCs'
obsp: 'connectivities', 'distances'
Calculating Diffusion Pseudotime using scanpy. This can be replaced by other methods, which can be implemented in your own preprocessing steps.
[4]:
adata1.uns['iroot'] = np.flatnonzero(adata1.obs['leiden'] == '4')[0]
sc.tl.dpt(adata1)
Now, pseudotime is stored in adata.obs
[5]:
adata1.obs
[5]:
| n_genes | leiden | dpt_pseudotime | |
|---|---|---|---|
| index | |||
| Run4_120703408880541 | 1697 | 1 | 0.660055 |
| Run4_120703409056541 | 1097 | 7 | 0.103425 |
| Run4_120703409580963 | 834 | 8 | 0.614602 |
| Run4_120703423990708 | 994 | 8 | 0.609378 |
| Run4_120703424252854 | 2277 | 19 | 0.507636 |
| ... | ... | ... | ... |
| Run5_241114589051630 | 2802 | 11 | 0.922035 |
| Run5_241114589051819 | 1695 | 10 | 0.474332 |
| Run5_241114589128940 | 2131 | 5 | 0.510828 |
| Run5_241114589357942 | 2982 | 11 | 0.920328 |
| Run5_241114589841822 | 2374 | 19 | 0.473308 |
5780 rows × 3 columns
[6]:
sc.pl.umap(adata1, color=["leiden", "dpt_pseudotime"])

Apply the same process to one or more datasets that you would like to align.
[7]:
adata2 = cp.dataset.velten17()
The dataset already exist in ../data/capital_dataset/velten17_capital.h5ad.
Reading datasets from ../data/capital_dataset/velten17_capital.h5ad.
[8]:
adata2.uns['iroot'] = np.flatnonzero(adata2.obs['leiden'] == '0')[0]
sc.tl.dpt(adata2)
[9]:
sc.pl.umap(adata2, color=["leiden", "dpt_pseudotime"])

Aligning trajectory trees¶
[10]:
cp.tl.trajectory_tree(adata1, root_node="4", groupby="leiden", tree=None)
[11]:
cp.tl.trajectory_tree(adata2, root_node="0", groupby="leiden", tree=None)
[12]:
cdata = cp.tl.tree_alignment(adata1, adata2, num_genes1=2000, num_genes2=2000)
Calculating tree alignment
411 genes are used to calculate cost of tree alignment.
Calculation finished.
Draw the tree alignment result.
[13]:
cp.pl.tree_alignment(cdata)

Applying preprocessed pseudotime for dynamic time warping¶
Specify the name of the column which stores pseudotime and the preprocessed pseudotime will be used in downstream analysis.
[14]:
cp.tl.dtw(cdata, gene=cdata.genes_for_tree_align, multi_genes=True, pseudotime="dpt_pseudotime")
[15]:
cp.pl.dtw(cdata, gene=["multi_genes"], alignment=["alignment001", "alignment006"], col_pseudotime="dpt_pseudotime",
data1_name="Setty+2019", data2_name="Velten+2017")

[16]:
main_markers = [
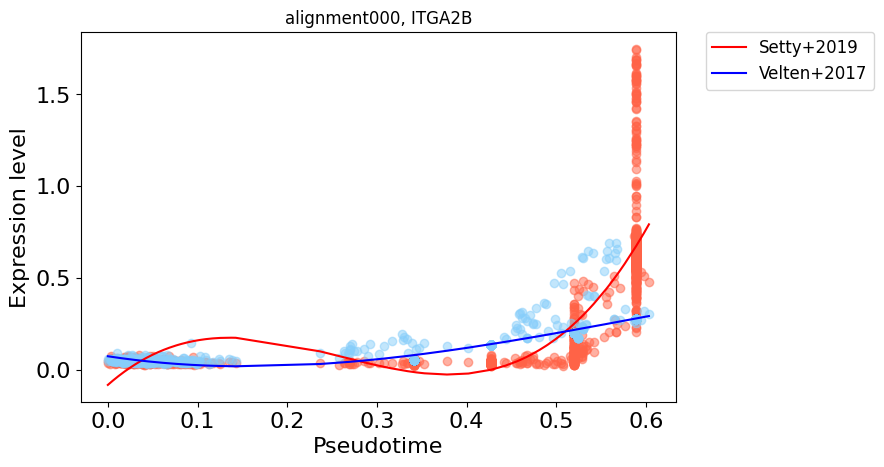
["alignment000", "ITGA2B"],
["alignment006", "LGMN"]
]
[17]:
for alignment, gene in main_markers:
cp.pl.gene_expression_trend(
cdata, gene=gene, alignment=alignment, fontsize=16, ticksize=16, multi_genes=True, col_pseudotime="dpt_pseudotime",
switch_psedotime=True,data1_name="Setty+2019", data2_name="Velten+2017", polyfit_dimension=3
)

[ ]: